Hi all,
Since the data continues to be downloaded at a fairly high rate, and the cost of fast bandwidth through Git-LFS was getting kind of high, I’ve moved the image files to a static location off of GitHub. I think this makes sense because they’re much larger than the label files when compressed, and they’re unlikely to change so a version history for them doesn’t add much value. The segmentation labels will remain on GitHub indefinitely. A python script to download the imaging from this new location and place it in the repository file tree like before has been added to the repository’s starter code. It’s designed to be robust to spotty internet connection. Please let me know if you have any questions or concerns with the new workflow.
Cheers!
Nick

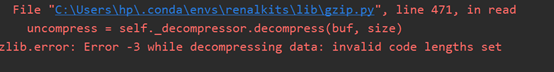 .
.